
Key Takeaways
- Experimental antibiotics disrupt a vital protein degradation system in Mycobacterium tuberculosis
- Findings explain distinct mechanisms of ecumicin, ilamycin, and cyclomarin
- Study highlights ClpC1–ClpP1P2 as a promising drug target for drug-resistant TB
A New Angle in Tuberculosis Drug Research
Experimental antibiotics targeting tuberculosis are gaining renewed focus following new mechanistic insights from researchers at the University of Sydney and the Centenary Institute. Published in Nature Communications, the study addresses a critical challenge in global health: the urgent need for new treatments against drug-resistant tuberculosis (TB).
TB continues to cause approximately 1.2 million deaths annually, remaining one of the deadliest infectious diseases worldwide. Resistance to existing therapies, particularly in the Asia-Pacific region, has intensified the demand for alternative antibacterial strategies.
How Do These Experimental Antibiotics Work on Tuberculosis (TB)?
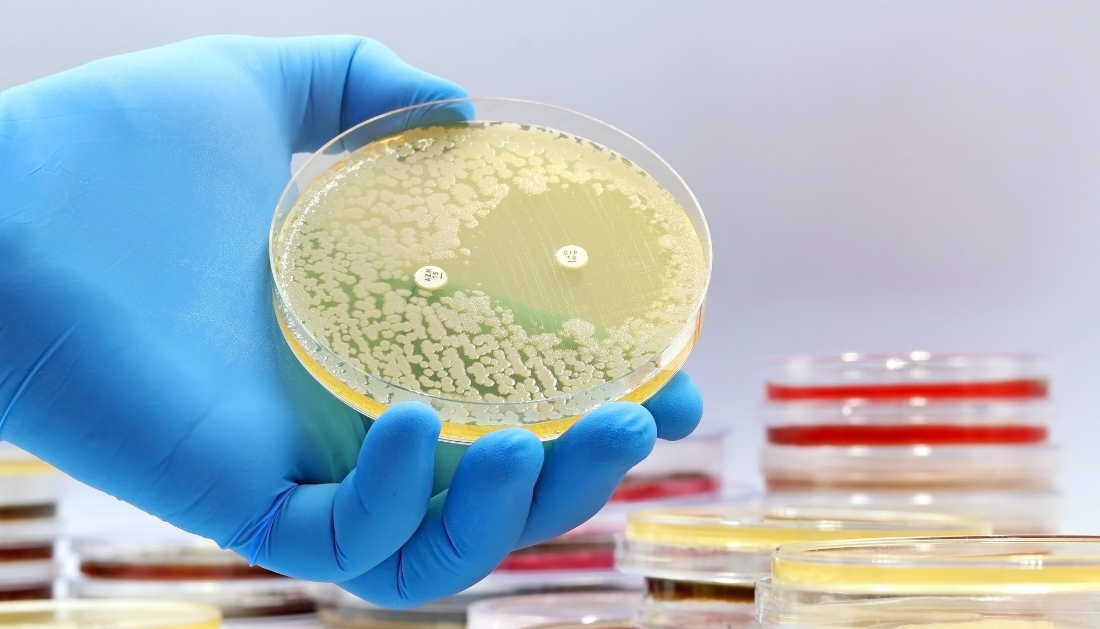
The research focused on three naturally derived antibiotic compounds, ecumicin, ilamycin, and cyclomarin, and their effects on a crucial bacterial system known as the ClpC1–ClpP1P2 protein degradation complex. This molecular machinery enables Mycobacterium tuberculosis to remove damaged or excess proteins, a function essential for bacterial survival under stress conditions such as host immune pressure.
Rather than simply halting this system, the study revealed that each compound disrupts the complex differently. Using proteomic analysis of more than 3,000 bacterial proteins, researchers demonstrated that interference with this single system leads to widespread protein imbalance across the bacterium.
Implications for Drug-Resistant TB Treatment
According to the investigators, these disruptions significantly weaken bacterial function and survival. For clinicians and infectious disease specialists, this insight is critical. It suggests that targeting bacterial protein quality-control systems may offer a viable therapeutic route, especially for multidrug-resistant TB cases where treatment options are limited.
The ClpC1–ClpP1P2 complex has been relatively underexplored as a drug target. Understanding how structurally distinct compounds interact with it provides a foundation for designing more precise anti-TB agents with improved efficacy.
Explore All Infectious Disease CME Conferences & Online Courses 2026
What This Means for Clinical Practice and Research
This study strengthens the pipeline for next-generation TB therapies by offering mechanistic clarity that can guide medicinal chemistry and translational research.
For healthcare professionals, it underscores the importance of continued innovation in antimicrobial development and highlights protein degradation pathways as a strategic focus in TB drug discovery.
Source:
more recommended stories
 Early Autism Detection Using Wearable Sensors in Infants
Early Autism Detection Using Wearable Sensors in InfantsKey Highlights: Wearable movement sensors may.
 Alzheimer’s Disease Driven by Cancer Gene Mutations
Alzheimer’s Disease Driven by Cancer Gene MutationsKey Highlights Microglial cells in Alzheimer’s.
 Cardiovascular Risk Rises with Heatwaves Over 38°C
Cardiovascular Risk Rises with Heatwaves Over 38°CKey Highlights Extreme heat (>38°C) significantly.
 Psychedelic Use Linked to Major Life Changes Study
Psychedelic Use Linked to Major Life Changes StudyQuick Summary 83% of users reported.
 Nutrition Research: Why Context Matters for Healthy Foods
Nutrition Research: Why Context Matters for Healthy FoodsQuick Summary Foods do not have.
 Hyperemesis Gravidarum Genes: Largest GWAS Study
Hyperemesis Gravidarum Genes: Largest GWAS StudyQuick Summary Largest GWAS identifies 10.
 Brain Categorization Redefined by New Neuroscience Study
Brain Categorization Redefined by New Neuroscience StudyKey Highlights Brain Categorization is not.
 GLP-1 Drug Effectiveness Reduced by Genetic Variants
GLP-1 Drug Effectiveness Reduced by Genetic VariantsKey Points Summary Genetic variants in.
 TB Vaccines Show Safety but Limited Protection in Large Trial
TB Vaccines Show Safety but Limited Protection in Large TrialKey Highlights Two candidate Tuberculosis vaccines,.
 Drug Overdose Spike 2025 Was a Modeling Error
Drug Overdose Spike 2025 Was a Modeling ErrorQuick Summary A reported U.S. drug.

Leave a Comment